Building a Benthic Genome Database for Improved Oil Spill Monitoring
– FEBRUARY 2, 2017
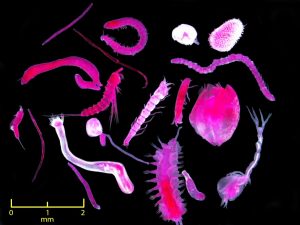
Meiofauna are invertebrate organisms that live in seafloor sediments. These marine creatures perform ecosystem functions such as trophic transfer, biogeochemical cycles, pollution removal, and sediment transport stability. Sensitive to environmental events such as oil spills, meiofauna are valuable bioindicators of impacts from contamination. However, their small size and our limited knowledge about these organisms’ community structure and taxonomy makes it difficult to track and characterize oil’s impacts.
The Gulf of Mexico Research Initiative recently awarded Dr. Kelley Thomas a grant to compile a reference genome database of benthic meiofaunal eukaryotes and establish Standard Operating Procedures (SOPs) for monitoring oil spills using bioinformatics. These tools can help researchers and responders interpret how spills might affect organisms in a particular area and how the organisms are expected to recover. “There’s so much we don’t understand about the relationship between the Gulf’s biogeochemical structure and the distribution of different taxa – we really don’t have any idea,” explained Thomas. “This information will help us better interpret the spill’s consequences and what kind of recovery will take place.”
His team is analyzing as many individual benthic eukaryotes as they can using whole genome shotgun sequencing, a quick and economically-effective method for analyzing the structure of long DNA strands. The researchers are compiling the resulting data into a large database of reference genomes to identify biomarker genes based on their ability to identify changes in population and community structure. Once completed, they can sequence pre- and post-spill samples of the benthic community for the biomarker genes and identify which organisms and communities the spill impacted.
“In the mid-2000s, genetic sequencing technology rapidly advanced to allow us to run up to 200,000,000 sequences at a time when before we could only run about 100 sequences at a time,” explained Thomas. “The technology has steadily improved and gotten cheaper over time since then. That’s why today it is feasible to consider modeling the benthic Gulf of Mexico in a large-scale way just by conducting DNA sequencing and analyses.”
The project is also developing clear and concise SOPs for using the genome database to conduct environmental and oil spill monitoring. The open-source SOPs will target researchers without previous genomics experience to help train future scientists to understand the communities’ sediment biology and conduct big data analyses using bioinformatics. An annual workshop will introduce undergraduate and graduate students to the SOPs, from sample preparation to interpreting metagenomics and implementing bioinformatics.
“It’s not just microscopes anymore. Most of a researcher’s time is spent in front of a computer,” said Thomas. “The skills taught in this training will have more biological and environmental applications than just learning how to analyze Gulf of Mexico data.”
Thomas’s team is enthusiastic about the potential of their project’s products to offer analytic tools and facilitate academic progress in the field of biology. They hope to fill gaps in the current understanding of benthic eukaryote taxa, including information about their roles in the environment, their interactions, and even their viral histories. He explained that his team is very interested in the possibility of combining their work with that of other GoMRI projects. For example, combining their findings with the ongoing effort to describe the Gulf’s biogeochemistry and other such patterns and trying to link it to organismal patterns.
“We’re not just developing a practical resource. I believe that this database is going to contribute to and make a substantial impact on biology,” said Thomas. “This is going to be the way people will monitor those environments.”
The project’s researchers are W. Kelley Thomas of the University of New Hampshire Hubbard Center for Genome Studies, Holly M. Bik of the University of California – Riverside, and Paul A. Montagna of the Texas A&M University – Corpus Christi Harte Research Institute for Gulf of Mexico Studies. Their project is Genomic Responses to the Deepwater Horizon Event and Development of High-Throughput Biological Assays for Oil Spills.
************
The Gulf of Mexico Research Initiative (GoMRI) is a 10-year independent research program established to study the effect, and the potential associated impact, of hydrocarbon releases on the environment and public health, as well as to develop improved spill mitigation, oil detection, characterization and remediation technologies. An independent and academic 20-member Research Board makes the funding and research direction decisions to ensure the intellectual quality, effectiveness and academic independence of the GoMRI research. All research data, findings and publications will be made publicly available. The program was established through a $500 million financial commitment from BP. For more information, visit https://gulfresearchinitiative.org/.
© Copyright 2010-2017 Gulf of Mexico Research Initiative (GoMRI) – All Rights Reserved. Redistribution is encouraged with acknowledgement to the Gulf of Mexico Research Initiative (GoMRI). Please credit images and/or videos as done in each article. Questions? Contact web-content editor Nilde “Maggie” Dannreuther, Northern Gulf Institute, Mississippi State University (maggied@ngi.msstate.edu).